Exploring the Mechanism Behind Drug Eruptions in the Skin
May 16, 2024/ Chiba University: Although drug eruptions are often linked to the human leukocyte antigen (HLA), the mechanism of its involvement in presenting symptoms of the skin remains unclear. In a recent study, researchers from Chiba University used genetically engineered mice to demonstrate the role of HLA in mediating intracellular reactions in keratinocytes, leading to drug eruptions in the skin. Their findings could lead to improved preventive and treatment measures for drug eruptions.
Although medications can often help patients find a cure or respite from their condition, millions of people worldwide suffer from unpredictable drug toxicities every year. In particular, drug eruptions which manifest through symptoms such as redness, blisters, and itching on the skin, are quite common. Severe drug eruptions can become life-threatening and can have long-lasting consequences. Thus, understanding how and why drug eruptions occur is an important area of research in medical science.
To this end, previous studies have identified specific variants of certain genes as potential causal agents of drug eruptions. Scientists believe that the genes encoding the human leukocyte antigen (HLA), a protein expressed on the surface of leucocytes known to play an important role in the immune system, are involved in the onset of drug eruption. However, current theories cannot explain why HLA-related drug eruptions typically manifest on the skin rather than in multiple organs throughout the body.
To address this knowledge gap, a research team including Lecturer Shigeki Aoki, Kousei Ito, and Akira Kazaoka from the Graduate School of Medical and Pharmaceutical Sciences, Chiba University, conducted an in-depth study on the link between HLA and drug eruptions. Their findings were published in PNAS Nexus on April 2, 2024.
The researchers first conducted a series of experiments on keratinocytes from mice, which are the primary type of cells found in the skin. These keratinocytes were engineered to express a specific variant of the HLA gene called HLA-B*57:01, which specifically bind to the antiviral drug abacavir. Then, they validated these findings in genetically modified mice expressing HLA-B*57:01, that were exposed to abacavir.
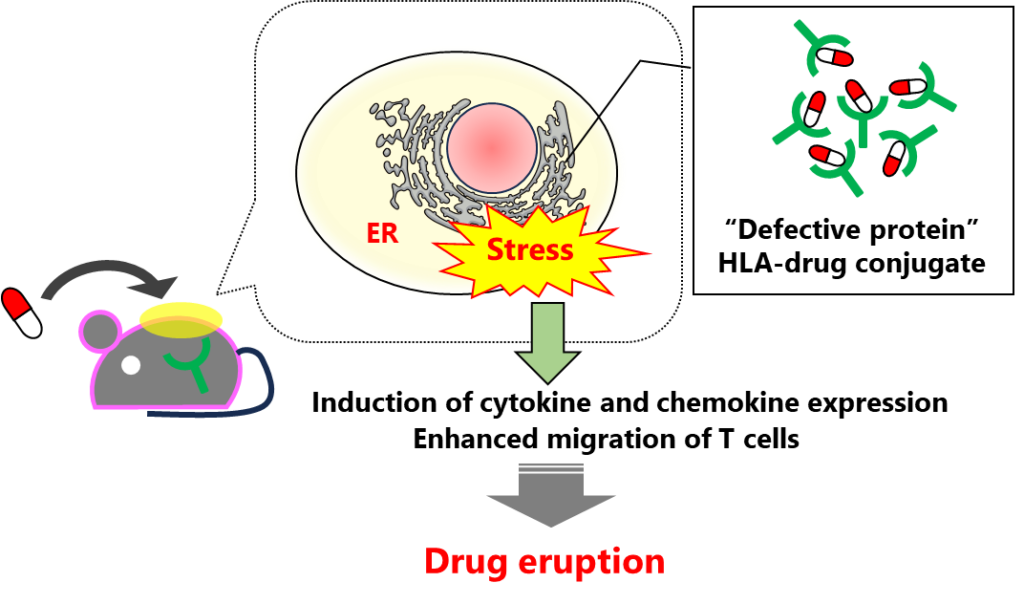
The researchers found that HLA-B*57:01 expressing keratinocytes that were exposed to abacavir exhibited endoplasmic reticulum (ER) stress responses, such as immediate release of calcium into the cytosol and elevated expression of heat shock protein 70 (HSP70). They also observed an increased production of cytokines and immune cell migration. Abacavir exposure triggered HLA misfolding in the ER, leading to ER stress. Moreover, the researchers observed that the ER stress could be reduced by using 4-phenylbutyrate (4-PB). By alleviating this stress, they managed to suppress the onset of severe drug eruption symptoms. This newfound knowledge could form the basis for innovative treatment options for management of drug eruptions.
But how does this new information contrast with what was already known about HLA? “HLA molecules are an integral component of our immune system, that typically present foreign antigens to white blood cells, which judge these antigens as self or non-self. In this established role, HLAs are usually secondary players,” explains Dr. Aoki. “However, our research highlights a novel function of the HLA molecule within skin cells. We revealed that a specific HLA genotype in keratinocytes can recognize certain drugs as foreign, triggering an endoplasmic reticulum stress response.”
Taken together, the findings of this study uncover a new role of HLA proteins in sensing and responding to potential threats in skin cells. Thus, their functions may extend well beyond mere antigen presentation for the immune system. Moreover, considering that the variant of HLA possessed by an individual can be determined through genetic testing, this study could help develop preventive measures and diagnostics against severe adverse drug reactions. According to Dr. Aoki, this is in line with current research directions and trends in medical science. “In 10 years, we anticipate entering the ‘whole genome era,’ where personalized medicine based on individual genomes will become a standard practice,” he comments. He further adds, “Building on the findings of this study, we believe that a comprehensive understanding of the mechanism underlying HLA-dependent adverse drug reactions will enable the delivery of safe medical care, allowing patients to avoid unnecessary suffering due to side effects.”
Overall, future investigations in this research area might minimize the occurrence of drug eruptions and save people from potentially fatal adverse drug reactions.
About Dr. Shigeki Aoki
Dr. Shigeki Aoki is a lecturer at the Graduate School of Pharmaceutical Sciences, Chiba University, Japan. His research focuses mainly on cancer metabolism and drug toxicity. He has authored multiple papers published in reputed journals. Dr. Aoki is a member of various professional bodies in Japan. He has also received several awards for his research, including the Award for Young Scientists conferred by The Pharmaceutical Society of Japan.
Funding:
This work was supported by the Japan Society for the Promotion of Science (JSPS) under JSPS KAKENHI Grant Nos. JP15H04661, JP15K14995, JP16K18932, JP17J03861, JP19H03386, JP21H02640, JP22J12718, JP23H02644, and JP23H04764.
Reference:
Title of original paper: HLA-B*57:01-dependent intracellular stress in keratinocytes triggers dermal hypersensitivity reactions to abacavir
Authors: Akira Kazaokaa, Sota Fujimoria, Yushiro Yamadaa, Tomohiro Shirayanagia, Yuying Gaoa, Saki Kuwaharaa, Naoki Sakamotoa, Takeshi Susukidaa,b, Shigeki Aokia, and Kousei Itoa
Affiliations:
aLaboratory of Biopharmaceutics, Graduate School of Pharmaceutical Sciences, Chiba University
bLaboratory of Cancer Biology and Immunology, Section of Host Defences, Institute of Natural Medicine, University of Toyama
Journal: PNAS Nexus
